
We're excited to share that Nanome has joined the OpenFold Consortium, a non-profit AI research consortium building open-source tools for biology and drug discovery.
OpenFold is home to some of the most impactful open-source work in structural biology right now. Their flagship, OpenFold3, is a fully open biomolecular structure prediction model developed by Mohammed AlQuraishi's lab at Columbia University and a growing community of industry and academic contributors. It's licensed under Apache 2.0, which means researchers and companies can test, retrain, fine-tune, and build on top of it freely. Other available co-folding methods, by comparison, restrict commercial use through lack of training code, training data, or even inference code.
The consortium just released a major OpenFold3 update alongside publicly available training datasets, making the entire training and inference stack reproducible and auditable by anyone. That kind of radical openness is exactly the ethos we believe in at Nanome.
Why This Matters to Us
At Nanome, we build the interface layer for scientific AI. Our platform spans immersive 3D molecular visualization in XR and our AI co-pilot MARA sits right at the point where computational predictions meet human scientific judgment. Structure prediction models like OpenFold3 generate the molecular geometries that researchers then need to see, interpret, manipulate, and act on.
"The gap between generating a predicted structure and actually using it in a drug discovery workflow is still massive," said Steve McCloskey, CEO and Co-Founder of Nanome. "OpenFold is doing the hard work of making structure prediction open and reproducible. We want to make sure the output of that work doesn't just live in a terminal somewhere. It needs to be in scientists' hands, in 3D, where they can make real decisions with it."
Joining OpenFold puts us at the table with organizations pushing the boundaries of what's computationally possible in drug discovery, while we focus on making those outputs usable — in context, in 3D, and alongside your team.
What It Looks Like in Practice

This is KRAS G12C — probably the most famous "undruggable" target in oncology — predicted by OpenFold3 and loaded into Nanome right alongside its experimental crystal structure. For context: KRAS burned through two decades of failed drug programs before Sotorasib finally cracked it in 2021. Here, a sequence goes in, a predicted geometry comes out, and within minutes your team is standing inside the structure together, comparing prediction against experiment, poking around the binding pocket, making actual decisions. No crystallography lab, no months of sample prep — just structure prediction to spatial collaboration in one sitting.
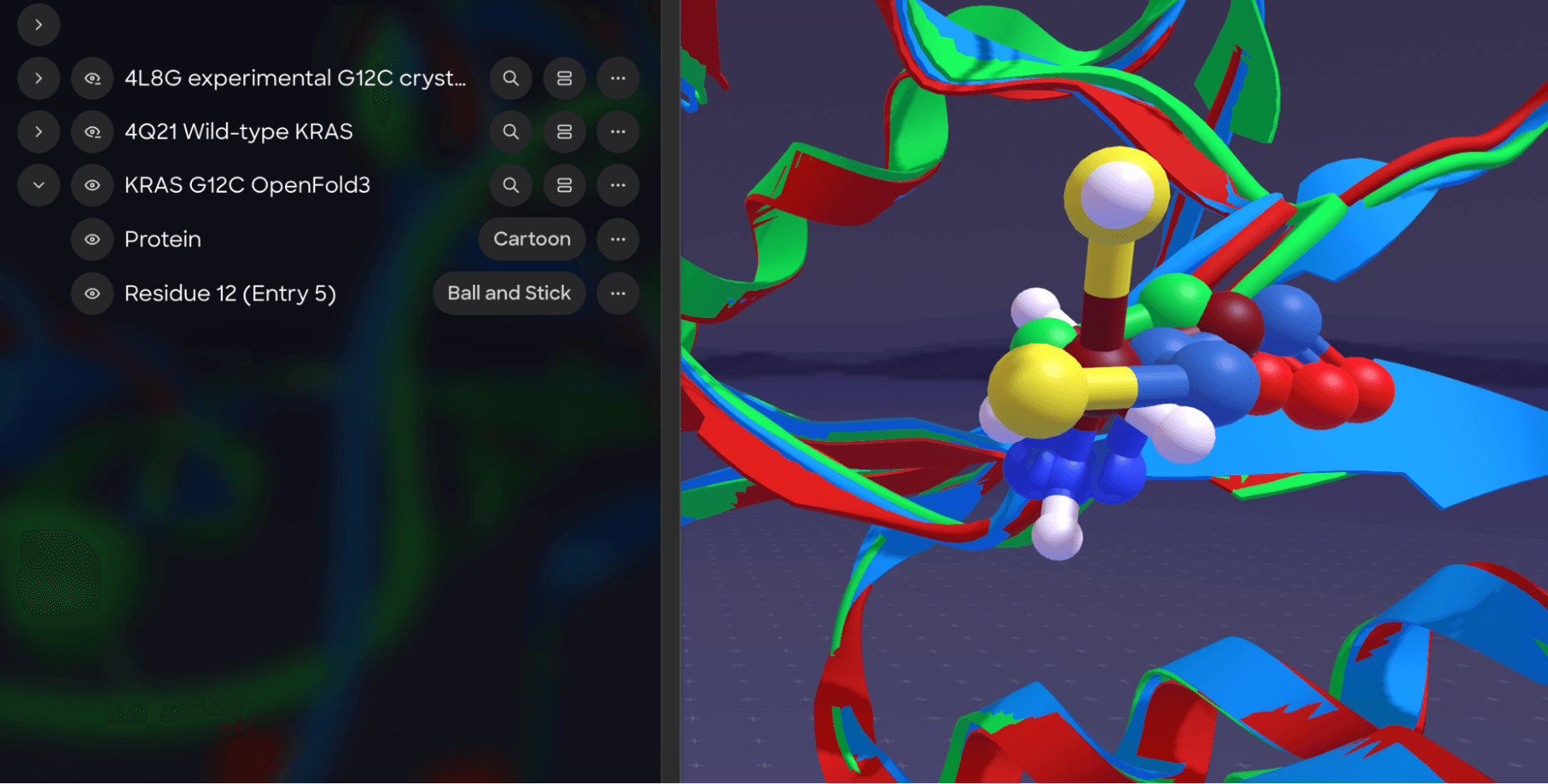

With our next release, this same view will be accessible directly from the web:
Who's in the Consortium

We're joining alongside global pharmaceutical leaders like Johnson & Johnson, Novo Nordisk, Roche, and Bristol Myers Squibb, as well as cutting-edge biotechs and tech companies like Cyrus Biotechnology, Structure Therapeutics, SandboxAQ, Lambda, Dassault Systèmes, and Outpace Bio. The full roster spans 30+ organizations across pharma, biotech, synthetic biology, GPU cloud, and scientific software, all contributing to the shared goal of making foundational AI for biology accessible to everyone.
What's Next
We're looking forward to contributing to the consortium's mission and exploring how Nanome's visualization and AI orchestration capabilities can complement the structural prediction tools coming out of OpenFold. As these models get more powerful, the need for intuitive, spatial interfaces to make sense of their outputs only grows.
More to come. If you're interested in how Nanome fits into the evolving AI drug discovery stack, reach out to us or follow along on our blog.
Nanome is the interface layer for scientific AI, combining immersive molecular visualization with MARA, our AI-powered scientific co-pilot. We serve enterprise pharmaceutical clients and research teams worldwide. Learn more at nanome.ai.